Figures and data in The unfolded protein response and endoplasmic reticulum protein targeting machineries converge on the stress sensor IRE1 | eLife
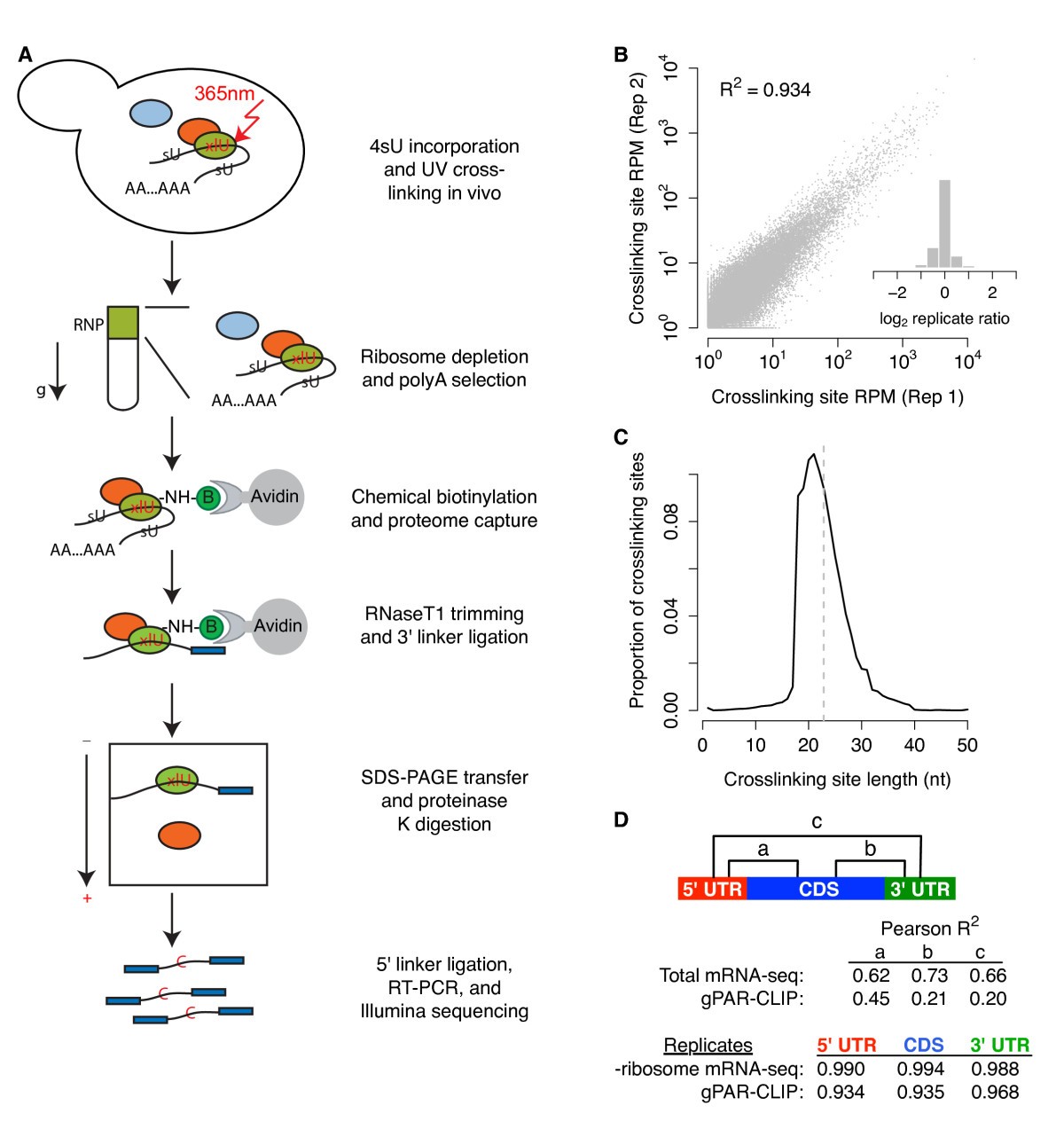
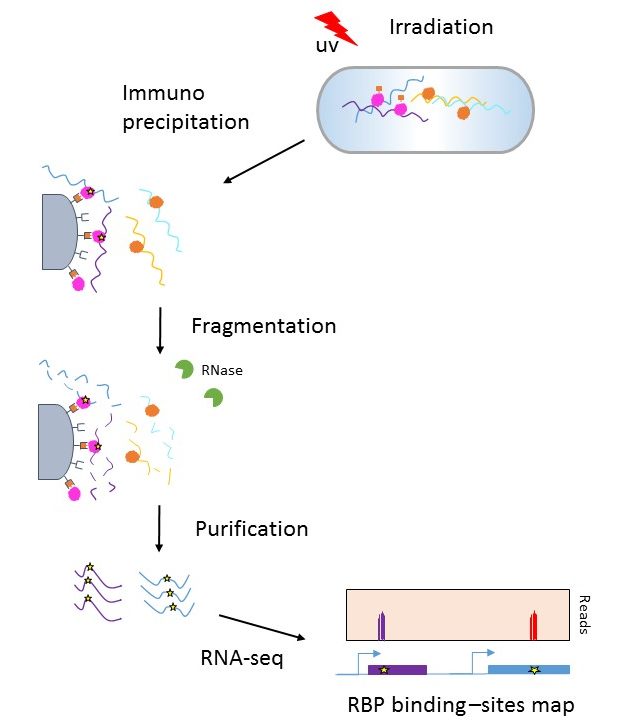
Pervasive and dynamic protein binding sites of the mRNA transcriptome in Saccharomyces cerevisiae | Genome Biology | Full Text
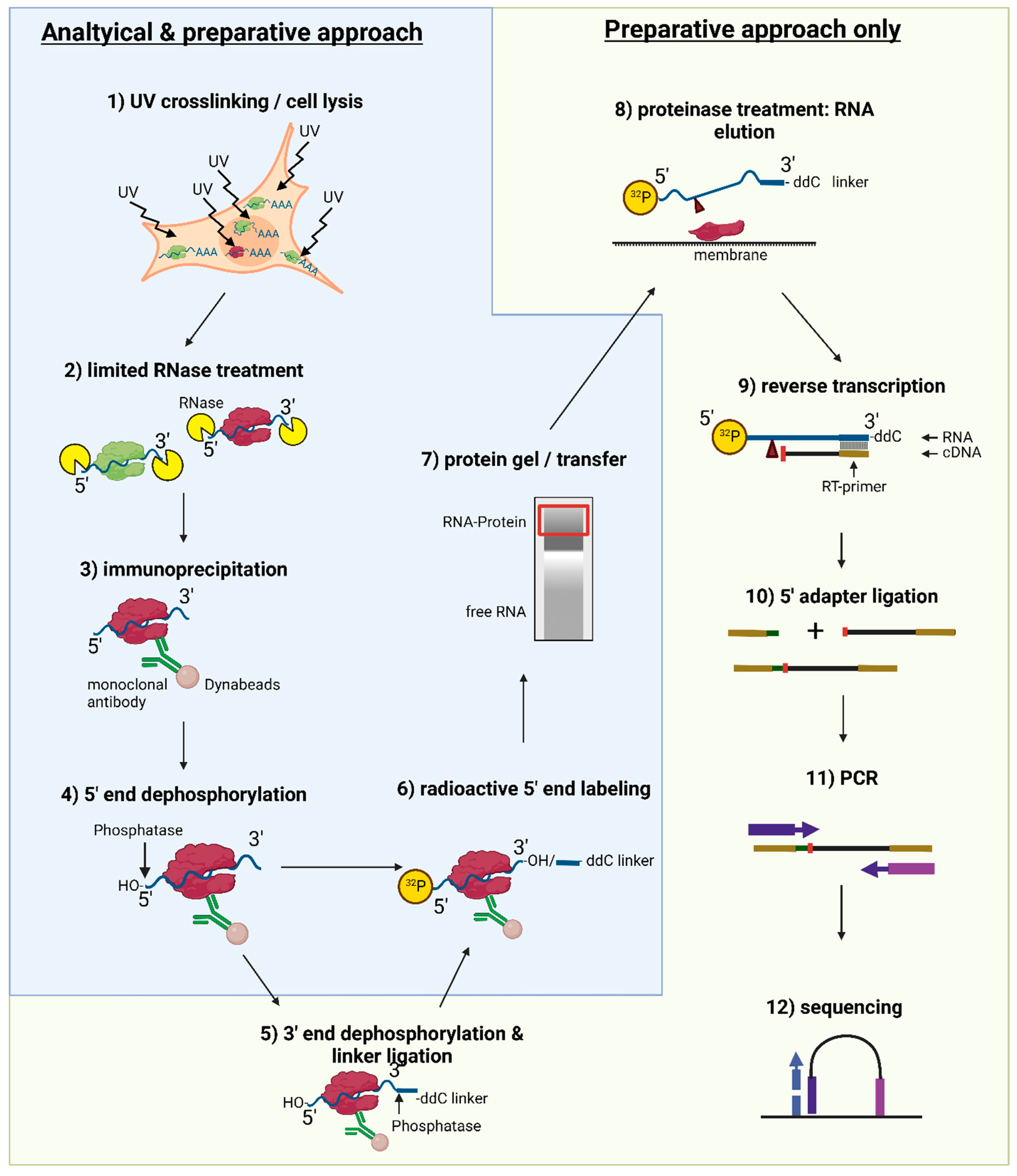
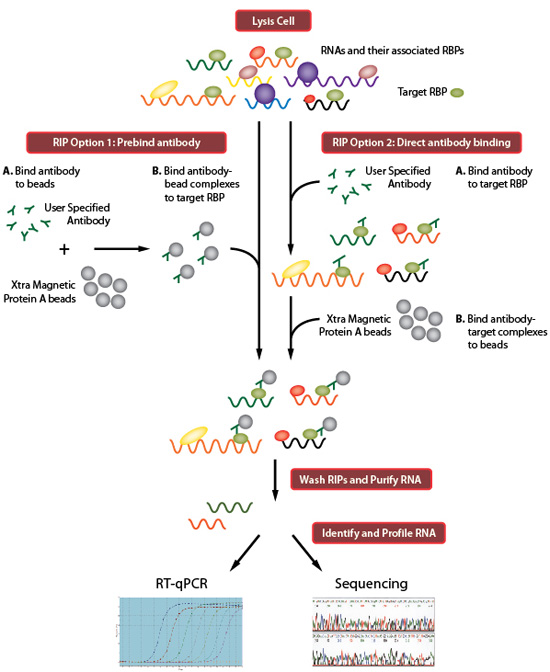
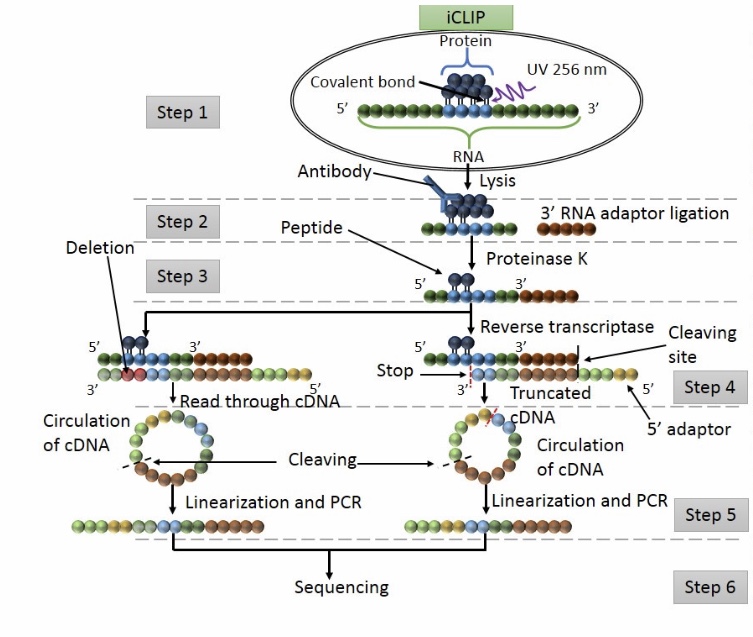
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein

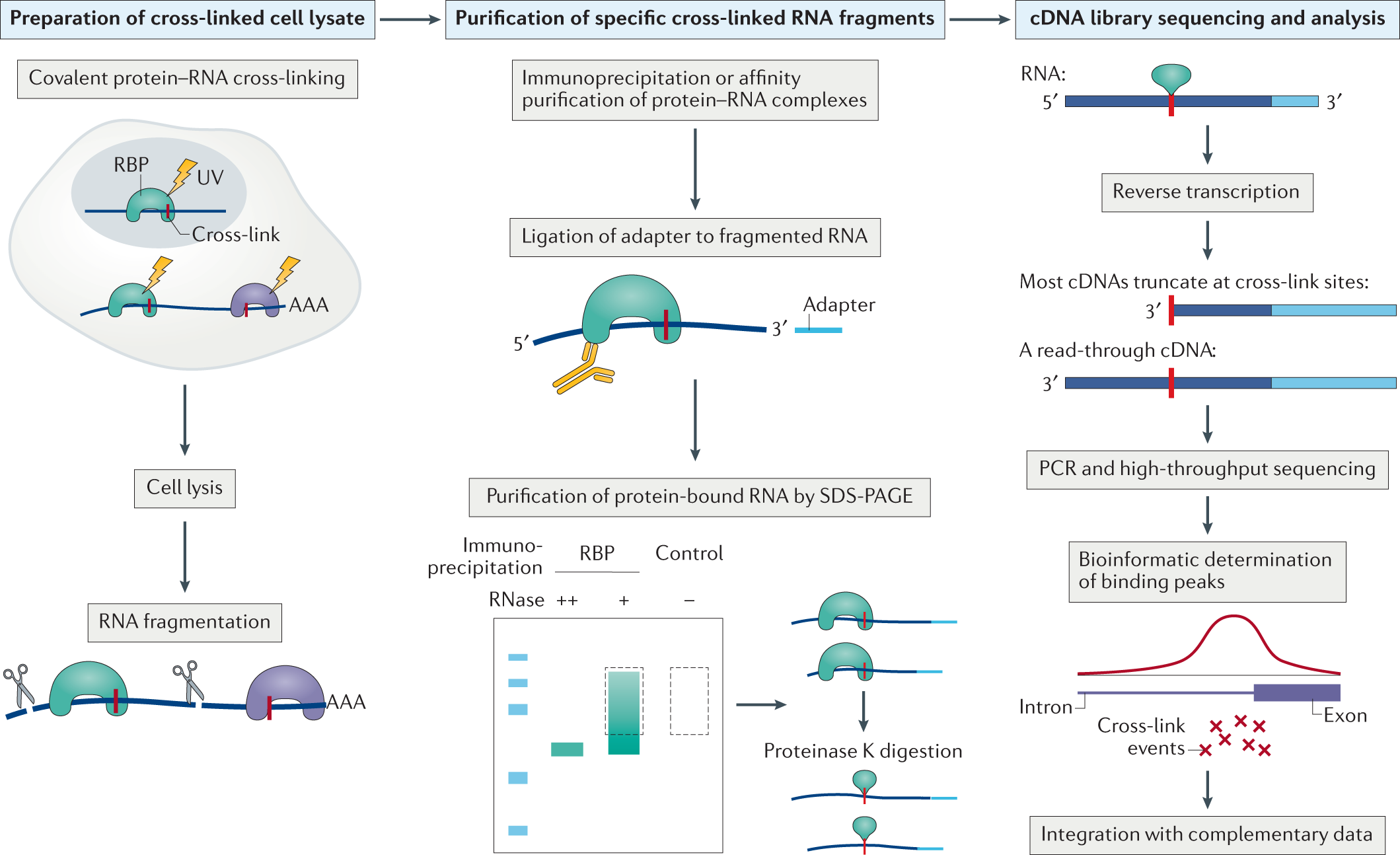
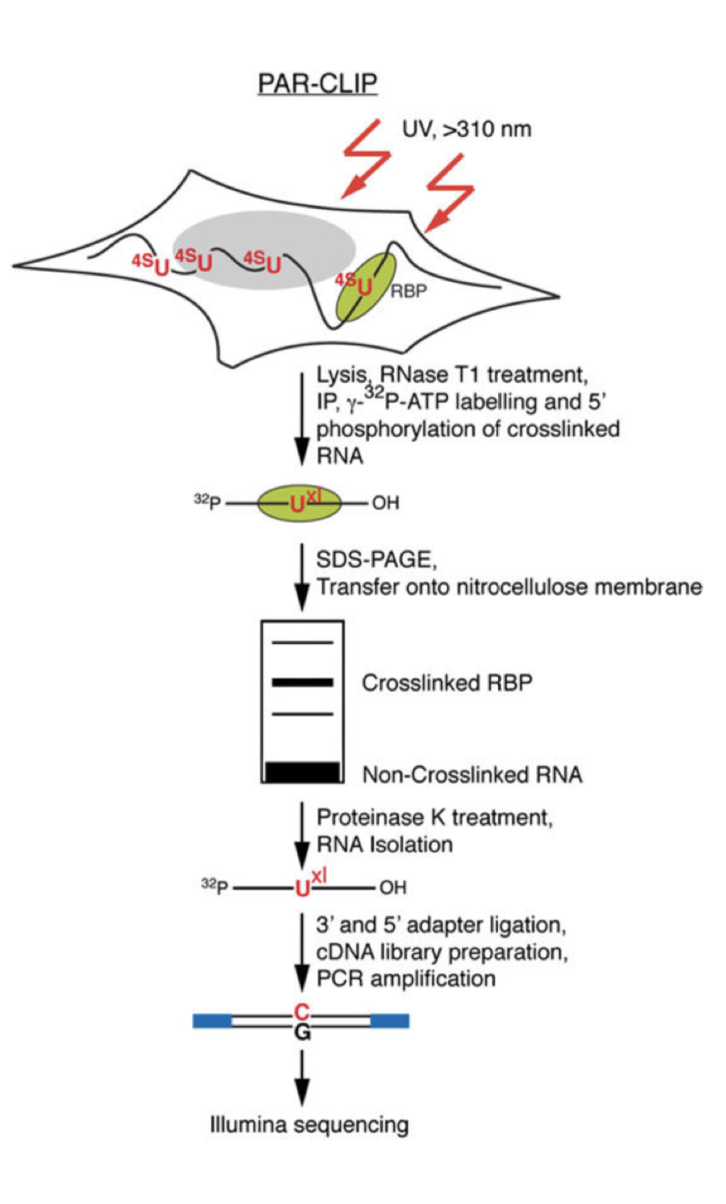
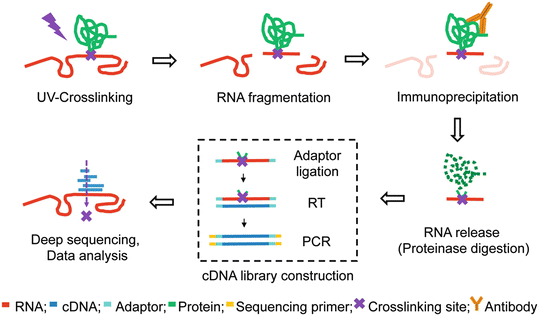
Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram
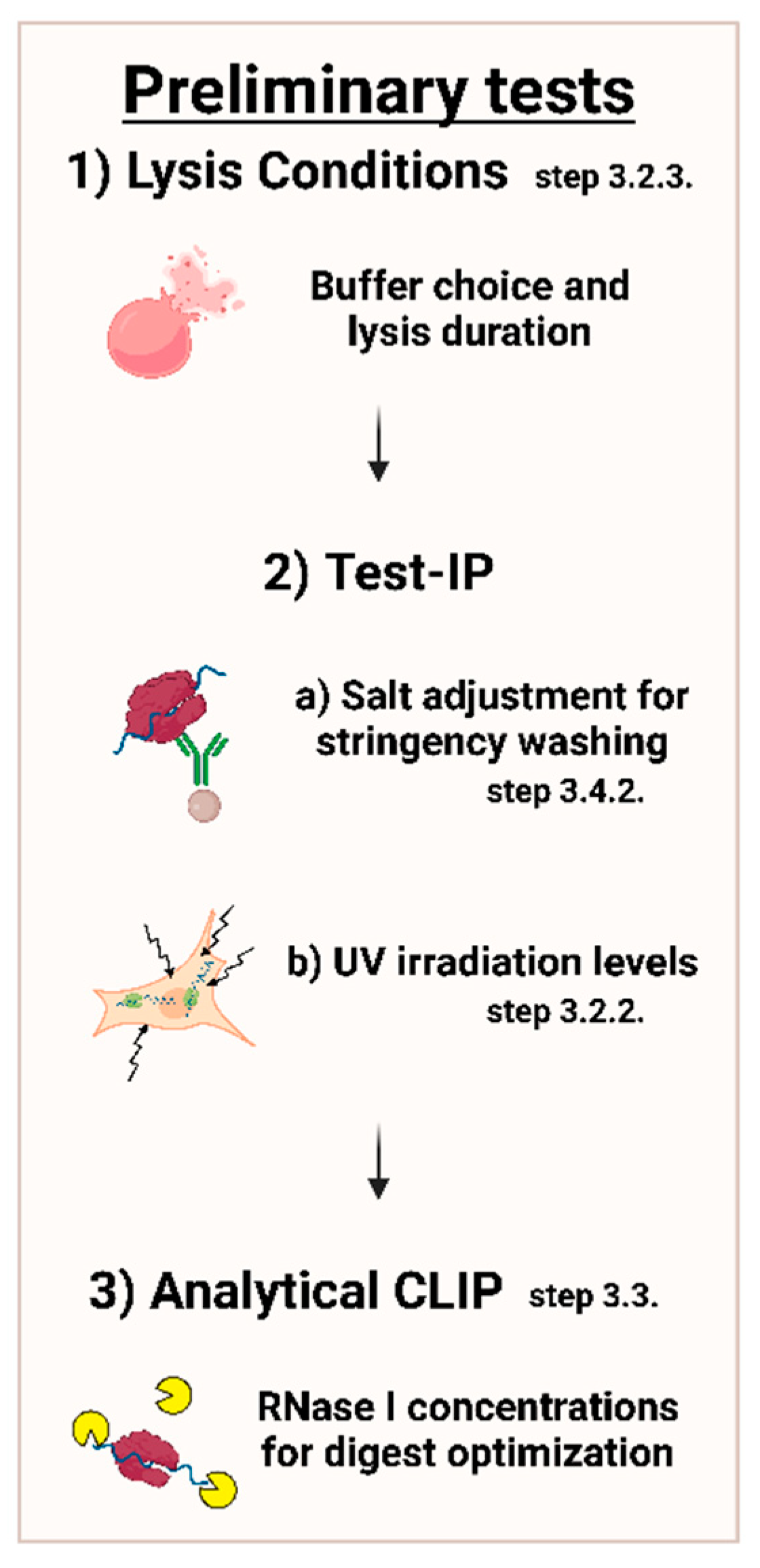
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein

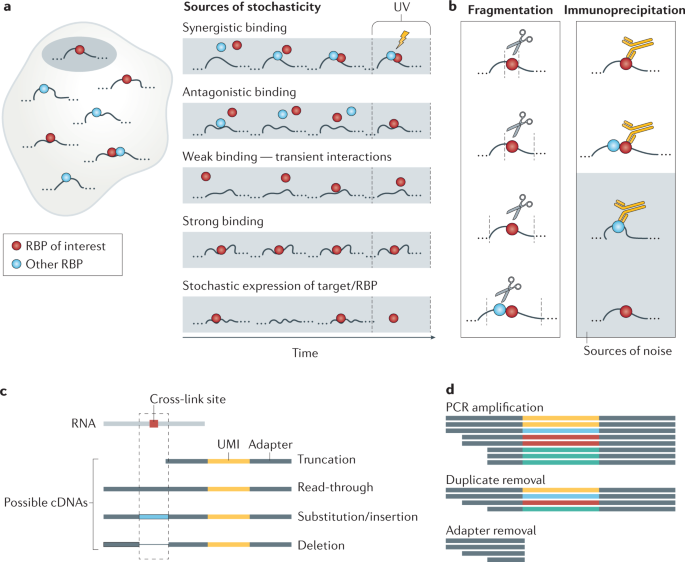
Mapping Argonaute and conventional RNA-binding protein interactions with RNA at single-nucleotide resolution using HITS-CLIP and CIMS analysis | Nature Protocols
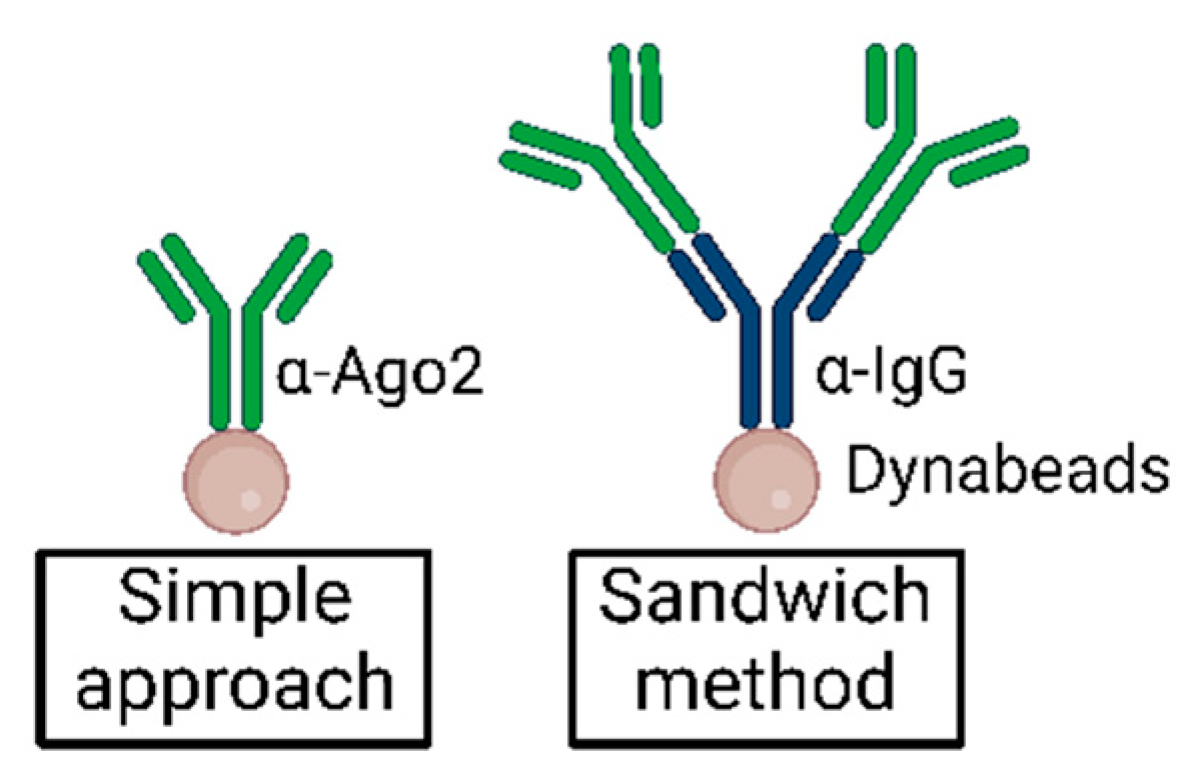


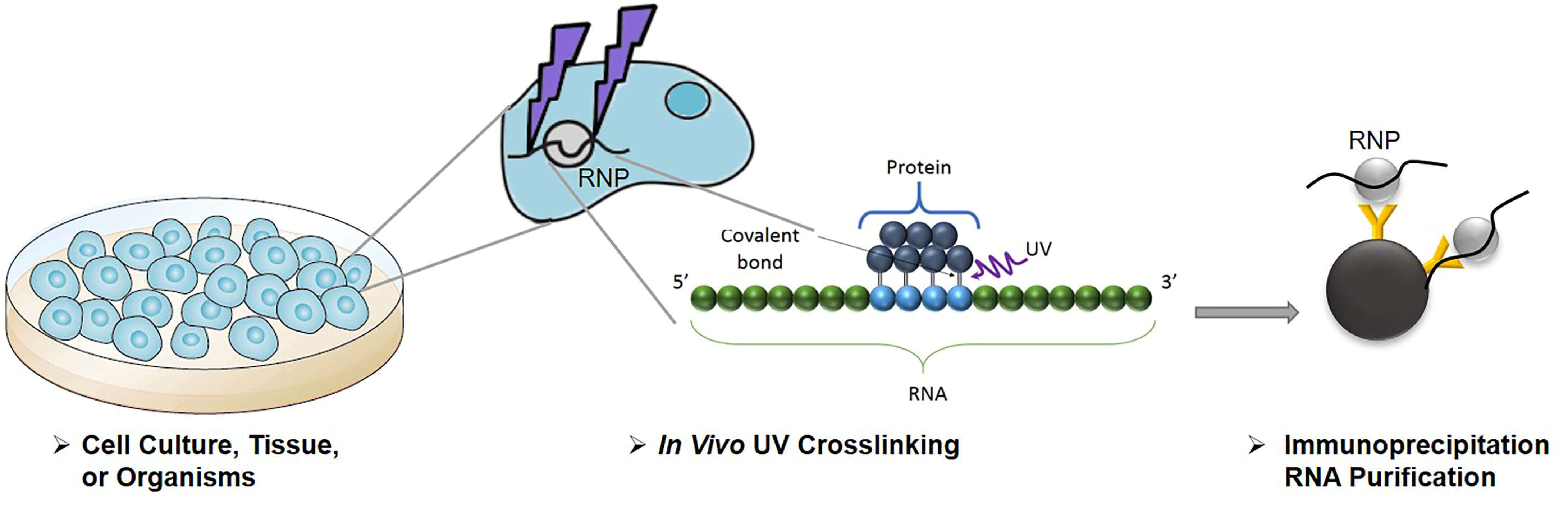





-2.jpg)